Stepwise and Hierarchical
Making the intercept and slopes makes sense!
- When to use depends on your questions. However, centering is safest
to do (and is often recommended)
- Centering
- Zscore
- POMP
- You need to decide on whether it makes sense to transform both DV and IVs or one or the other.
- Let’s make a practice dataset to explore
- We will transform just the IVs for now:
library(car) #graph data
library(stargazer)
# IQ scores of 5 people
Y<-c(85, 90, 100, 120, 140)
# Likert scale rating of liking of reading books (1 hate to 7 love)
X1<-c(1,2,4,6,7)
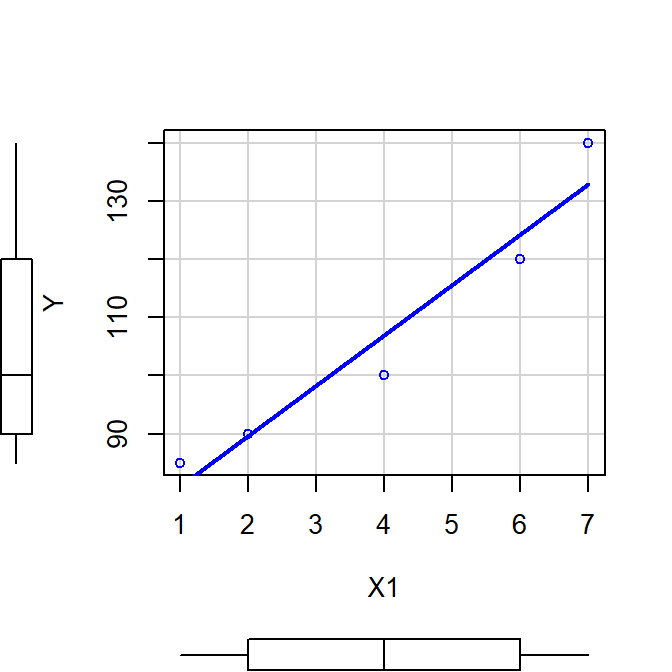
scatterplot(Y~X1, smooth=FALSE)
Mr<-lm(Y~X1)
stargazer(Mr,type="html",
intercept.bottom = FALSE, notes.append = FALSE, header=FALSE)| Dependent variable: | |
| Y | |
| Constant | 72.385*** |
| (6.010) | |
| X1 | 8.654*** |
| (1.305) | |
| Observations | 5 |
| R2 | 0.936 |
| Adjusted R2 | 0.915 |
| Residual Std. Error | 6.655 (df = 3) |
| F Statistic | 43.958*** (df = 1; 3) |
| Note: | p<0.1; p<0.05; p<0.01 |
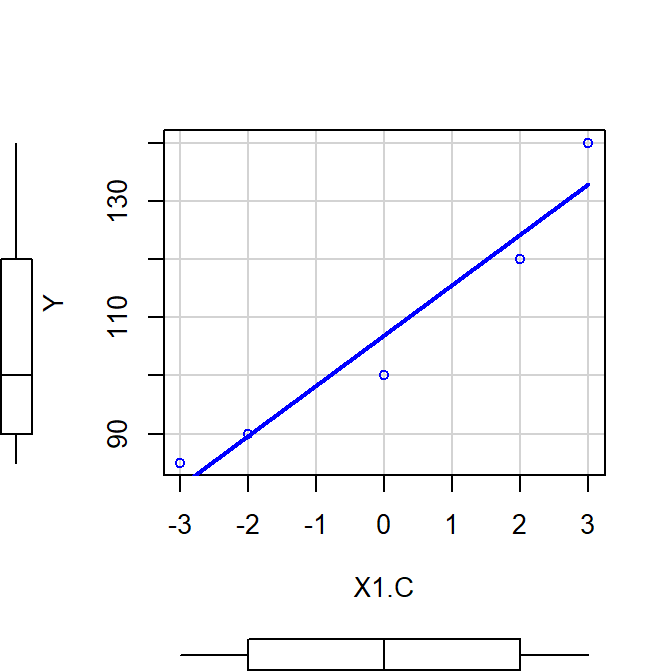
Center
- \(Center = {X - M}\)
- Intercept is not at the MEAN of IV (no 0 of IV)
- Does NOT changes meaning of slope
- R:
scale(Data,scale=FALSE)[,]- scale add a dimension to our new variable, and we can remove it
using [,]
- We usually don’t need this, but it can mess up sometime down the road
- scale add a dimension to our new variable, and we can remove it
using [,]
X1.C<-scale(X1,scale=FALSE)[,]
scatterplot(Y~X1.C, smooth=FALSE)
Mc<-lm(Y~X1.C)
stargazer(Mc,type="html",
intercept.bottom = FALSE, notes.append = FALSE, header=FALSE)| Dependent variable: | |
| Y | |
| Constant | 107.000*** |
| (2.976) | |
| X1.C | 8.654*** |
| (1.305) | |
| Observations | 5 |
| R2 | 0.936 |
| Adjusted R2 | 0.915 |
| Residual Std. Error | 6.655 (df = 3) |
| F Statistic | 43.958*** (df = 1; 3) |
| Note: | p<0.1; p<0.05; p<0.01 |
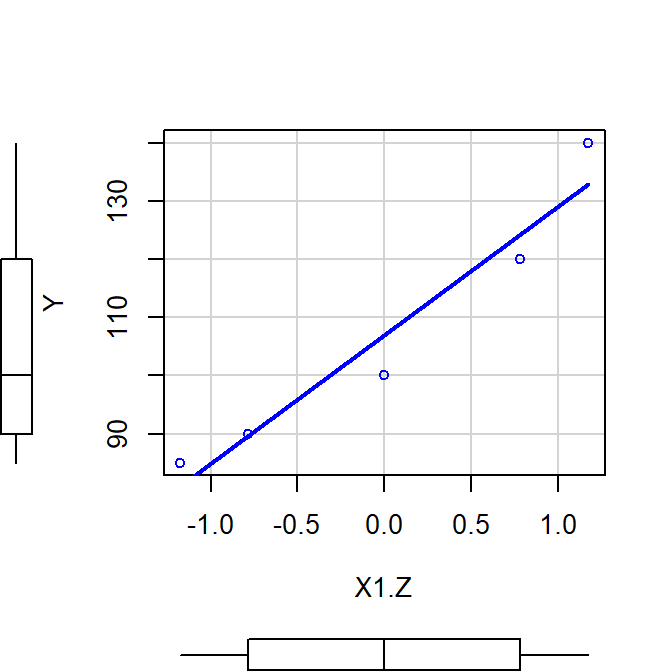
Zscore
- \(Z = \frac{X - M}{s}\)
- Intercept is not at the MEAN of IV (no 0 of IV)
- Slope changes meaning: no longer in unites of original DV, now in sd units
- R:
scale(data)[,]
#Zscore
X1.Z<-scale(X1)[,]
scatterplot(Y~X1.Z, smooth=FALSE)
Mz<-lm(Y~X1.Z)
stargazer(Mz,type="html",
intercept.bottom = FALSE, notes.append = FALSE, header=FALSE)| Dependent variable: | |
| Y | |
| Constant | 107.000*** |
| (2.976) | |
| X1.Z | 22.063*** |
| (3.328) | |
| Observations | 5 |
| R2 | 0.936 |
| Adjusted R2 | 0.915 |
| Residual Std. Error | 6.655 (df = 3) |
| F Statistic | 43.958*** (df = 1; 3) |
| Note: | p<0.1; p<0.05; p<0.01 |
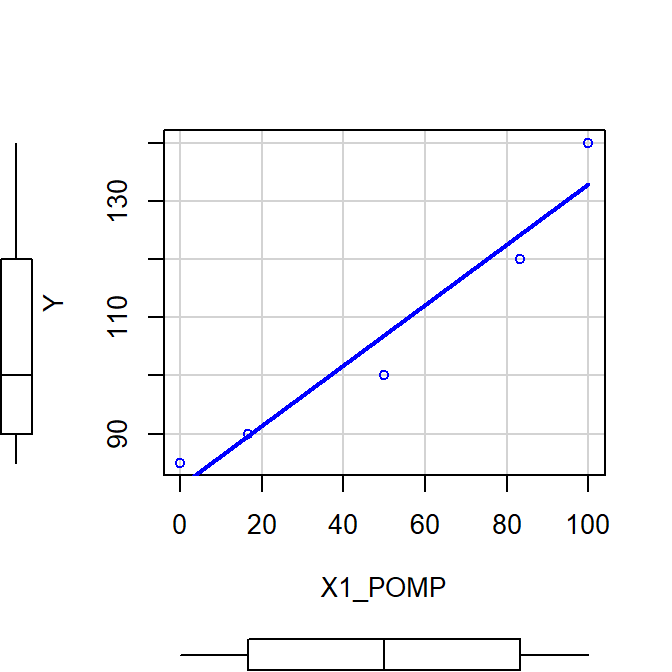
POMP
- \(POMP = \frac{X - MinX}{Max_X - Min_X}*100\)
- Note: I like to X 100 cause I find it easier to think in percent (not proportion)
- Useful when data are bounded (or scaled funny)
- Intercept is again at 0 of IV [but the slopes is different, so the intercept changes a bit]
- Does changes meaning of slope: is now a function of percent change of IV
X1_POMP = (X1 - min(X1)) / (max(X1) - min(X1))*100
scatterplot(Y~X1_POMP, smooth=FALSE)
Mp<-lm(Y~X1_POMP)
stargazer(Mp,type="html",
intercept.bottom = FALSE, notes.append = FALSE, header=FALSE)| Dependent variable: | |
| Y | |
| Constant | 81.038*** |
| (4.919) | |
| X1_POMP | 0.519*** |
| (0.078) | |
| Observations | 5 |
| R2 | 0.936 |
| Adjusted R2 | 0.915 |
| Residual Std. Error | 6.655 (df = 3) |
| F Statistic | 43.958*** (df = 1; 3) |
| Note: | p<0.1; p<0.05; p<0.01 |
Simultaneous Regression (standard approach)
- Put all your variables in and see what the effect is of each term
- Very conservative approach
- Does not allow you to understand additive effects very easily
- You noticed this problem when we were trying to explain Health ~ Years married + Age
- Had you only looked at this final model you might never have understood that Years married acted as a good predictor on its own.
- Also what if you have a theory you want to test? You need to see the additive effects.
Hierarchical Modeling
- Is the change in \(R^2\), meaningful (Model 2 \(R^2\) - Model 1 \(R^2\))?
- The order in which models are run are meaningful
- Terms in models do not need to be analyzed one at a time, but can be entered as ‘sets’
- a set of variables are theoretically or experimentally driven
- So Model 2 \(R^2\) - Model 1 \(R^2\) meaningful?
Hierarchical Modeling driven by the researcher
- Forward selection: Start with simple models and get more complex nested models
- Backward selection: Start with complex nested models and get more simple
- Stepwise selection: can be viewed as a variation of the forward selection method (one predictor at a time) but predictors are deleted in subsequent steps if they no longer contribute appreciable unique prediction
- Which you choose is can depend on how you like to ask questions
Forward Selection of nested models
- A common approach “model building”
- Again let’s make up our dummy data
library(MASS) #create data
py1 =.6 #Cor between X1 (ice cream) and happiness
py2 =.4 #Cor between X2 (Brownies) and happiness
p12= .2 #Cor between X1 (ice cream) and X2 (Brownies)
Means.X1X2Y<- c(10,10,10) #set the means of X and Y variables
CovMatrix.X1X2Y <- matrix(c(1,p12,py1, p12,1,py2, py1,py2,1),3,3) # creates the covariate matrix
set.seed(42)
CorrDataT<-mvrnorm(n=100, mu=Means.X1X2Y,Sigma=CovMatrix.X1X2Y, empirical=TRUE)
CorrDataT<-as.data.frame(CorrDataT)
colnames(CorrDataT) <- c("IceCream","Brownies","Happiness")library(corrplot)
corrplot(cor(CorrDataT), method = "number")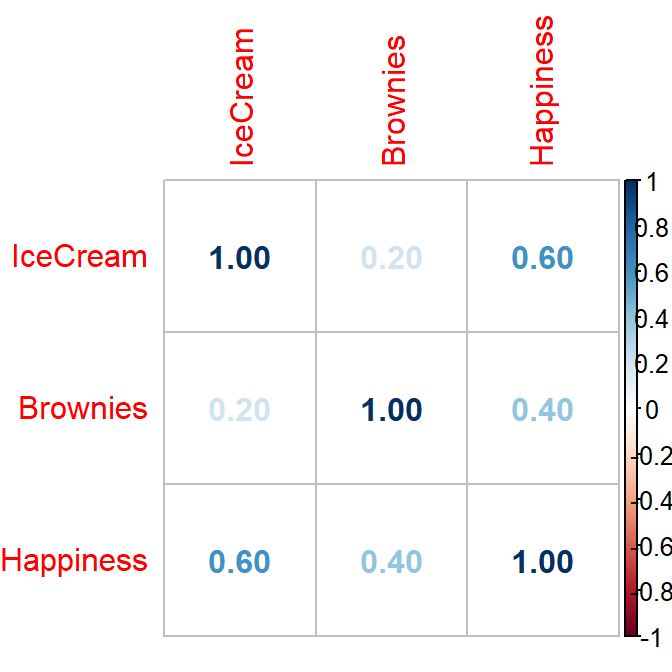
First alittle side track…
- Remember the \(R2\) values are reported as F values right?
- This means you can actually get an ANOVA like table for the model
- for example:
###############Model 1
Ice.Model<-lm(Happiness~ IceCream, data = CorrDataT)
anova(Ice.Model)## Analysis of Variance Table
##
## Response: Happiness
## Df Sum Sq Mean Sq F value Pr(>F)
## IceCream 1 35.64 35.640 55.125 4.193e-11 ***
## Residuals 98 63.36 0.647
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1- The \(R2\) this is explained to unexplained variance (like in our ANOVA)
- \(R^2 = \frac{SS_{explained}}{SS_{explained}+SS_{residual}}\)
- just to check: anova(Ice.Model) 64.36
- which matched the \(R^2\) that R gives us 0.36
- When we check to see which model is best we actually test the differences
Lets forward-fit our models
- Model 1 (Smaller model)
Ice.Model<-lm(Happiness~ IceCream, data = CorrDataT)
R2.Model.1<-summary(Ice.Model)$r.squared- Model 2 (Larger model)
###############Model 1
Ice.Brown.Model<-lm(Happiness~ IceCream+Brownies, data = CorrDataT)
R2.Model.2<-summary(Ice.Brown.Model)$r.squaredlibrary(stargazer)
stargazer(Ice.Model,Ice.Brown.Model,type="html",
column.labels = c("Model 1", "Model 2"),
intercept.bottom = FALSE,
single.row=FALSE,
star.cutoffs = c(0.1, 0.05, 0.01, 0.001),
star.char = c("@", "*", "**", "***"),
notes= c("@p < .1 *p < .05 **p < .01 ***p < .001"),
notes.append = FALSE, header=FALSE)| Dependent variable: | ||
| Happiness | ||
| Model 1 | Model 2 | |
| (1) | (2) | |
| Constant | 4.000*** | 1.667@ |
| (0.812) | (0.982) | |
| IceCream | 0.600*** | 0.542*** |
| (0.081) | (0.077) | |
| Brownies | 0.292*** | |
| (0.077) | ||
| Observations | 100 | 100 |
| R2 | 0.360 | 0.442 |
| Adjusted R2 | 0.353 | 0.430 |
| Residual Std. Error | 0.804 (df = 98) | 0.755 (df = 97) |
| F Statistic | 55.125*** (df = 1; 98) | 38.366*** (df = 2; 97) |
| Note: | @p < .1 p < .05 p < .01 p < .001 | |
- Let’s the difference in \(R^2\)
- \(R_{Change}^2\) =\(R_{Larger}^2\) - \(R_{Smaller}^2\)
- In R, we call for function
anovaand use an \(F\) where the degrees of freedom is the number of parameter differences between Larger and Smaller model
R2.Change<-R2.Model.2-R2.Model.1
anova(Ice.Model,Ice.Brown.Model)## Analysis of Variance Table
##
## Model 1: Happiness ~ IceCream
## Model 2: Happiness ~ IceCream + Brownies
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 98 63.360
## 2 97 55.275 1 8.085 14.188 0.0002838 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1- The \(R_{Change}^2\) = 0.0816667 is
significant
- So, in other words, we see model 2 fit the data better than model 1.
Backward-fitting of nested models
- You as does taking away variables reduce my \(R^2\) significantly
- Sometimes used to validate you have a parsimonious model
- You might forward-fit a set of variables and backward fit critical ones to test a specific hypothesis
- Using the same data as above, we will get the same values (just
negative)
- \(R_{Change}^2\) =\(R_{smaller}^2\) - \(R_{Larger}^2\)
###############Model 1.B
Ice.Brown.Model<-lm(Happiness~ IceCream+Brownies, data = CorrDataT)
R2.Model.1.B<-summary(Ice.Brown.Model)$r.squared
###############Model 2.B
Ice.Model<-lm(Happiness~ IceCream, data = CorrDataT)
R2.Model.2.B<-summary(Ice.Model)$r.squared
R2.Change.B<-R2.Model.2.B-R2.Model.1.B
anova(Ice.Brown.Model,Ice.Model)## Analysis of Variance Table
##
## Model 1: Happiness ~ IceCream + Brownies
## Model 2: Happiness ~ IceCream
## Res.Df RSS Df Sum of Sq F Pr(>F)
## 1 97 55.275
## 2 98 63.360 -1 -8.085 14.188 0.0002838 ***
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1- The \(R_{Change}^2\) = -0.0816667
is significant
- So, in other words, we see model 1 is a worse fit of the data than model 2
Stepwise modeling by Computer
- Stepwise with many predicts is often done by computer and it does not always assume nested models (you can add and remove at the same)
- Exploratory: you have too many predictors and have no idea where to start
- You give the computer a larger number of predictors, and the computer decides the best fit model
- Sounds good, right? No, as the results can be unstable
- Change one variable in the set and the final model can change
- High chance of type I and type II error
- The computer makes decisions based on Akaike information criterion (AIC) not selected based on a change in \(R^2\), because models are not nested
- also computer makes decisions purely on fit values and has nothing do with a theory
- Solutions are often unique to that particular dataset
- The best model is often the one that parses a theory and only a human can do that at present
- Not really publishable because of these problems
Parsing influence
- As models get bigger and bigger its becomes a challenge to figure out the unique contribution to \(R^2\) of each variable
- There are many computation solutions that you can select from, but we will use one called lmg
- you can read about all the different ones here: https://core.ac.uk/download/pdf/6305006.pdf
- these methods are not well known in psychology, but can be very useful when people ask you what the relative importance of each variable is
- two approaches: show absolute \(R^2\) for each term or the relative % of \(R^2\) for each term
library(relaimpo)
# In terms of R2
calc.relimp(Ice.Brown.Model) ## Response variable: Happiness
## Total response variance: 1
## Analysis based on 100 observations
##
## 2 Regressors:
## IceCream Brownies
## Proportion of variance explained by model: 44.17%
## Metrics are not normalized (rela=FALSE).
##
## Relative importance metrics:
##
## lmg
## IceCream 0.3208333
## Brownies 0.1208333
##
## Average coefficients for different model sizes:
##
## 1X 2Xs
## IceCream 0.6 0.5416667
## Brownies 0.4 0.2916667# as % of R2
calc.relimp(Ice.Brown.Model,rela = TRUE) ## Response variable: Happiness
## Total response variance: 1
## Analysis based on 100 observations
##
## 2 Regressors:
## IceCream Brownies
## Proportion of variance explained by model: 44.17%
## Metrics are normalized to sum to 100% (rela=TRUE).
##
## Relative importance metrics:
##
## lmg
## IceCream 0.7264151
## Brownies 0.2735849
##
## Average coefficients for different model sizes:
##
## 1X 2Xs
## IceCream 0.6 0.5416667
## Brownies 0.4 0.2916667Final notes:
If you play with lots of predictors and do lots of models, something will be significant
Type I error is a big problem because of the ‘researcher degree of freedom problem’
Type II increases as a function of the number of predictors. a) you slice too much pie, b) each variable might try to each eat someone else’s slice
Less is more: ask targeted questions with as orthogonal a set of variables as you can
LS0tDQp0aXRsZTogIlN0ZXB3aXNlIGFuZCBIaWVyYXJjaGljYWwiDQpvdXRwdXQ6DQogIGh0bWxfZG9jdW1lbnQ6DQogICAgY29kZV9kb3dubG9hZDogeWVzDQogICAgZm9udHNpemU6IDhwdA0KICAgIGhpZ2hsaWdodDogdGV4dG1hdGUNCiAgICBudW1iZXJfc2VjdGlvbnM6IG5vDQogICAgdGhlbWU6IGZsYXRseQ0KICAgIHRvYzogeWVzDQogICAgdG9jX2Zsb2F0Og0KICAgICAgY29sbGFwc2VkOiBubw0KLS0tDQpgYGB7ciwgZWNobz1GQUxTRSwgd2FybmluZz1GQUxTRX0NCiNzZXR3ZCgnQzovVXNlcnMvQWxleFVJQy9Cb3ggU3luYy81NDUgUmVncmVzc2lvbiBTcHJpbmcgMjAxOC9XZWVrIDMgLSBNUicpDQojc2V0d2QoJ0M6L0FsZXhGaWxlcy9TdWdlclN5bmMvVUlDL1RlYWNoaW5nL0dyYWR1YXRlLzU0NS1TcHJpbmcyMDE4L1dlZWsgNSAtIFN0ZXAgYW5kIEhpZXJhcmNoaWNhbCcpDQpgYGANCg0KYGBge3Igc2V0dXAsIGluY2x1ZGU9RkFMU0V9DQojIHNldHVwIGZvciBSbm90ZWJvb2tzDQprbml0cjo6b3B0c19jaHVuayRzZXQoZWNobyA9IFRSVUUpICNTaG93IGFsbCBzY3JpcHQgYnkgZGVmYXVsdA0Ka25pdHI6Om9wdHNfY2h1bmskc2V0KG1lc3NhZ2UgPSBGQUxTRSkgI2hpZGUgbWVzc2FnZXMgDQprbml0cjo6b3B0c19jaHVuayRzZXQod2FybmluZyA9ICBGQUxTRSkgI2hpZGUgcGFja2FnZSB3YXJuaW5ncyANCmtuaXRyOjpvcHRzX2NodW5rJHNldChmaWcud2lkdGg9My41KSAjU2V0IGRlZmF1bHQgZmlndXJlIHNpemVzDQprbml0cjo6b3B0c19jaHVuayRzZXQoZmlnLmhlaWdodD0zLjUpICNTZXQgZGVmYXVsdCBmaWd1cmUgc2l6ZXMNCmtuaXRyOjpvcHRzX2NodW5rJHNldChmaWcuYWxpZ249J2NlbnRlcicpICNTZXQgZGVmYXVsdCBmaWd1cmUNCmtuaXRyOjpvcHRzX2NodW5rJHNldChmaWcuc2hvdyA9ICJob2xkIikgI1NldCBkZWZhdWx0IGZpZ3VyZQ0KYGBgDQoNClxwYWdlYnJlYWsNCg0KIyBNYWtpbmcgdGhlIGludGVyY2VwdCBhbmQgc2xvcGVzIG1ha2VzIHNlbnNlIQ0KLSBXaGVuIHRvIHVzZSBkZXBlbmRzIG9uIHlvdXIgcXVlc3Rpb25zLiBIb3dldmVyLCBjZW50ZXJpbmcgaXMgc2FmZXN0IHRvIGRvIChhbmQgaXMgb2Z0ZW4gcmVjb21tZW5kZWQpIA0KICAgIC0gQ2VudGVyaW5nIA0KICAgIC0gWnNjb3JlIA0KICAgIC0gUE9NUA0KLSBZb3UgbmVlZCB0byBkZWNpZGUgb24gd2hldGhlciBpdCBtYWtlcyBzZW5zZSB0byB0cmFuc2Zvcm0gYm90aCBEViBhbmQgSVZzIG9yIG9uZSBvciB0aGUgb3RoZXIuIA0KLSBMZXQncyBtYWtlIGEgcHJhY3RpY2UgZGF0YXNldCB0byBleHBsb3JlDQotIFdlIHdpbGwgdHJhbnNmb3JtIGp1c3QgdGhlIElWcyBmb3Igbm93OiANCg0KYGBge3IsIHJlc3VsdHM9J2FzaXMnfQ0KbGlicmFyeShjYXIpICNncmFwaCBkYXRhDQpsaWJyYXJ5KHN0YXJnYXplcikNCiMgSVEgc2NvcmVzIG9mIDUgcGVvcGxlDQpZPC1jKDg1LCA5MCwgMTAwLCAxMjAsIDE0MCkNCiMgTGlrZXJ0IHNjYWxlIHJhdGluZyBvZiBsaWtpbmcgb2YgcmVhZGluZyBib29rcyAoMSBoYXRlIHRvIDcgbG92ZSkNClgxPC1jKDEsMiw0LDYsNykNCnNjYXR0ZXJwbG90KFl+WDEsIHNtb290aD1GQUxTRSkNCk1yPC1sbShZflgxKQ0Kc3RhcmdhemVyKE1yLHR5cGU9Imh0bWwiLA0KICAgICAgICAgIGludGVyY2VwdC5ib3R0b20gPSBGQUxTRSwgbm90ZXMuYXBwZW5kID0gRkFMU0UsIGhlYWRlcj1GQUxTRSkNCmBgYA0KDQojIyBDZW50ZXINCi0gJENlbnRlciA9IHtYIC0gTX0kDQotIEludGVyY2VwdCBpcyBub3QgYXQgdGhlIE1FQU4gb2YgSVYgKG5vIDAgb2YgSVYpDQotIERvZXMgTk9UIGNoYW5nZXMgbWVhbmluZyBvZiBzbG9wZQ0KLSBSOiBgc2NhbGUoRGF0YSxzY2FsZT1GQUxTRSlbLF1gDQogICAgLSBzY2FsZSBhZGQgYSBkaW1lbnNpb24gdG8gb3VyIG5ldyB2YXJpYWJsZSwgYW5kIHdlIGNhbiByZW1vdmUgaXQgdXNpbmcgWyxdDQogICAgICAgIC0gV2UgdXN1YWxseSBkb24ndCBuZWVkIHRoaXMsIGJ1dCBpdCBjYW4gbWVzcyB1cCBzb21ldGltZSBkb3duIHRoZSByb2FkDQoNCmBgYHtyLCByZXN1bHRzPSdhc2lzJ30NClgxLkM8LXNjYWxlKFgxLHNjYWxlPUZBTFNFKVssXQ0Kc2NhdHRlcnBsb3QoWX5YMS5DLCBzbW9vdGg9RkFMU0UpDQpNYzwtbG0oWX5YMS5DKQ0Kc3RhcmdhemVyKE1jLHR5cGU9Imh0bWwiLA0KICAgICAgICAgIGludGVyY2VwdC5ib3R0b20gPSBGQUxTRSwgbm90ZXMuYXBwZW5kID0gRkFMU0UsIGhlYWRlcj1GQUxTRSkNCmBgYA0KDQojIyBac2NvcmUNCi0gJFogPSBcZnJhY3tYIC0gTX17c30kDQotIEludGVyY2VwdCBpcyBub3QgYXQgdGhlIE1FQU4gb2YgSVYgKG5vIDAgb2YgSVYpDQotIFNsb3BlIGNoYW5nZXMgbWVhbmluZzogbm8gbG9uZ2VyIGluIHVuaXRlcyBvZiBvcmlnaW5hbCBEViwgbm93IGluICpzZCogdW5pdHMNCi0gUjogYHNjYWxlKGRhdGEpWyxdYA0KDQpgYGB7ciwgcmVzdWx0cz0nYXNpcyd9DQojWnNjb3JlDQpYMS5aPC1zY2FsZShYMSlbLF0gDQpzY2F0dGVycGxvdChZflgxLlosIHNtb290aD1GQUxTRSkNCk16PC1sbShZflgxLlopDQpzdGFyZ2F6ZXIoTXosdHlwZT0iaHRtbCIsDQogICAgICAgICAgaW50ZXJjZXB0LmJvdHRvbSA9IEZBTFNFLCBub3Rlcy5hcHBlbmQgPSBGQUxTRSwgaGVhZGVyPUZBTFNFKQ0KYGBgDQoNCiMjIFBPTVANCi0gJFBPTVAgPSBcZnJhY3tYIC0gTWluWH17TWF4X1ggLSBNaW5fWH0qMTAwJA0KLSBOb3RlOiBJIGxpa2UgdG8gWCAxMDAgY2F1c2UgSSBmaW5kIGl0IGVhc2llciB0byB0aGluayBpbiBwZXJjZW50IChub3QgcHJvcG9ydGlvbikNCi0gVXNlZnVsIHdoZW4gZGF0YSBhcmUgYm91bmRlZCAob3Igc2NhbGVkIGZ1bm55KQ0KLSBJbnRlcmNlcHQgaXMgYWdhaW4gYXQgMCBvZiBJViBbYnV0IHRoZSBzbG9wZXMgaXMgZGlmZmVyZW50LCBzbyB0aGUgaW50ZXJjZXB0IGNoYW5nZXMgYSBiaXRdIA0KLSBEb2VzIGNoYW5nZXMgbWVhbmluZyBvZiBzbG9wZTogaXMgbm93IGEgZnVuY3Rpb24gb2YgcGVyY2VudCBjaGFuZ2Ugb2YgSVYgDQoNCmBgYHtyLCByZXN1bHRzPSdhc2lzJ30NClgxX1BPTVAgPSAoWDEgLSBtaW4oWDEpKSAvIChtYXgoWDEpIC0gbWluKFgxKSkqMTAwDQpzY2F0dGVycGxvdChZflgxX1BPTVAsIHNtb290aD1GQUxTRSkNCk1wPC1sbShZflgxX1BPTVApDQpzdGFyZ2F6ZXIoTXAsdHlwZT0iaHRtbCIsDQogICAgICAgICAgaW50ZXJjZXB0LmJvdHRvbSA9IEZBTFNFLCBub3Rlcy5hcHBlbmQgPSBGQUxTRSwgaGVhZGVyPUZBTFNFKQ0KYGBgDQoNClxwYWdlYnJlYWsNCg0KIyBTaW11bHRhbmVvdXMgUmVncmVzc2lvbiAoc3RhbmRhcmQgYXBwcm9hY2gpDQotIFB1dCBhbGwgeW91ciB2YXJpYWJsZXMgaW4gYW5kIHNlZSB3aGF0IHRoZSBlZmZlY3QgaXMgb2YgZWFjaCB0ZXJtDQotIFZlcnkgY29uc2VydmF0aXZlIGFwcHJvYWNoDQotIERvZXMgbm90IGFsbG93IHlvdSB0byB1bmRlcnN0YW5kIGFkZGl0aXZlIGVmZmVjdHMgdmVyeSBlYXNpbHkNCi0gWW91IG5vdGljZWQgdGhpcyBwcm9ibGVtIHdoZW4gd2Ugd2VyZSB0cnlpbmcgdG8gZXhwbGFpbiBIZWFsdGggfiBZZWFycyBtYXJyaWVkICsgQWdlDQotIEhhZCB5b3Ugb25seSBsb29rZWQgYXQgdGhpcyBmaW5hbCBtb2RlbCB5b3UgbWlnaHQgbmV2ZXIgaGF2ZSB1bmRlcnN0b29kIHRoYXQgWWVhcnMgbWFycmllZCBhY3RlZCBhcyBhIGdvb2QgcHJlZGljdG9yIG9uIGl0cyBvd24uIA0KLSBBbHNvIHdoYXQgaWYgeW91IGhhdmUgYSB0aGVvcnkgeW91IHdhbnQgdG8gdGVzdD8gWW91IG5lZWQgdG8gc2VlIHRoZSBhZGRpdGl2ZSBlZmZlY3RzLiANCg0KIyBIaWVyYXJjaGljYWwgTW9kZWxpbmcNCi0gSXMgdGhlIGNoYW5nZSBpbiAkUl4yJCwgbWVhbmluZ2Z1bCAoTW9kZWwgMiAkUl4yJCAtIE1vZGVsIDEgJFJeMiQpPw0KLSBUaGUgb3JkZXIgaW4gd2hpY2ggbW9kZWxzIGFyZSBydW4gYXJlIG1lYW5pbmdmdWwNCi0gVGVybXMgaW4gbW9kZWxzIGRvIG5vdCBuZWVkIHRvIGJlIGFuYWx5emVkIG9uZSBhdCBhIHRpbWUsIGJ1dCBjYW4gYmUgZW50ZXJlZCBhcyAnc2V0cycNCi0gYSBzZXQgb2YgdmFyaWFibGVzIGFyZSB0aGVvcmV0aWNhbGx5IG9yIGV4cGVyaW1lbnRhbGx5IGRyaXZlbiANCi0gU28gTW9kZWwgMiAkUl4yJCAtIE1vZGVsIDEgJFJeMiQgIG1lYW5pbmdmdWw/DQoNCiMjIEhpZXJhcmNoaWNhbCBNb2RlbGluZyBkcml2ZW4gYnkgdGhlIHJlc2VhcmNoZXINCi0gRm9yd2FyZCBzZWxlY3Rpb246IFN0YXJ0IHdpdGggc2ltcGxlIG1vZGVscyBhbmQgZ2V0IG1vcmUgY29tcGxleCBuZXN0ZWQgbW9kZWxzDQotIEJhY2t3YXJkIHNlbGVjdGlvbjogU3RhcnQgd2l0aCBjb21wbGV4IG5lc3RlZCBtb2RlbHMgYW5kIGdldCBtb3JlIHNpbXBsZQ0KLSBTdGVwd2lzZSBzZWxlY3Rpb246IGNhbiBiZSB2aWV3ZWQgYXMgYSB2YXJpYXRpb24gb2YgdGhlIGZvcndhcmQgc2VsZWN0aW9uIG1ldGhvZCAob25lIHByZWRpY3RvciBhdCBhIHRpbWUpIGJ1dCBwcmVkaWN0b3JzIGFyZSBkZWxldGVkIGluIHN1YnNlcXVlbnQgc3RlcHMgaWYgdGhleSBubyBsb25nZXIgY29udHJpYnV0ZSBhcHByZWNpYWJsZSB1bmlxdWUgcHJlZGljdGlvbg0KLSBXaGljaCB5b3UgY2hvb3NlIGlzIGNhbiBkZXBlbmQgb24gaG93IHlvdSBsaWtlIHRvIGFzayBxdWVzdGlvbnMNCg0KIyMjIEZvcndhcmQgU2VsZWN0aW9uIG9mIG5lc3RlZCBtb2RlbHMNCi0gQSBjb21tb24gYXBwcm9hY2ggIm1vZGVsIGJ1aWxkaW5nIg0KLSBBZ2FpbiBsZXQncyBtYWtlIHVwIG91ciBkdW1teSBkYXRhDQoNCmBgYHtyfQ0KbGlicmFyeShNQVNTKSAjY3JlYXRlIGRhdGENCnB5MSA9LjYgI0NvciBiZXR3ZWVuIFgxIChpY2UgY3JlYW0pIGFuZCBoYXBwaW5lc3MNCnB5MiA9LjQgI0NvciBiZXR3ZWVuIFgyIChCcm93bmllcykgYW5kIGhhcHBpbmVzcw0KcDEyPSAuMiAjQ29yIGJldHdlZW4gWDEgKGljZSBjcmVhbSkgYW5kIFgyIChCcm93bmllcykNCk1lYW5zLlgxWDJZPC0gYygxMCwxMCwxMCkgI3NldCB0aGUgbWVhbnMgb2YgWCBhbmQgWSB2YXJpYWJsZXMNCkNvdk1hdHJpeC5YMVgyWSA8LSBtYXRyaXgoYygxLHAxMixweTEsIHAxMiwxLHB5MiwgcHkxLHB5MiwxKSwzLDMpICMgY3JlYXRlcyB0aGUgY292YXJpYXRlIG1hdHJpeCANCnNldC5zZWVkKDQyKQ0KQ29yckRhdGFUPC1tdnJub3JtKG49MTAwLCBtdT1NZWFucy5YMVgyWSxTaWdtYT1Db3ZNYXRyaXguWDFYMlksIGVtcGlyaWNhbD1UUlVFKQ0KQ29yckRhdGFUPC1hcy5kYXRhLmZyYW1lKENvcnJEYXRhVCkNCmNvbG5hbWVzKENvcnJEYXRhVCkgPC0gYygiSWNlQ3JlYW0iLCJCcm93bmllcyIsIkhhcHBpbmVzcyIpDQpgYGANCg0KDQpgYGB7cn0NCmxpYnJhcnkoY29ycnBsb3QpDQpjb3JycGxvdChjb3IoQ29yckRhdGFUKSwgbWV0aG9kID0gIm51bWJlciIpDQpgYGANCg0KDQojIyMjIEZpcnN0IGFsaXR0bGUgc2lkZSB0cmFjay4uLg0KLSBSZW1lbWJlciB0aGUgJFIyJCB2YWx1ZXMgYXJlIHJlcG9ydGVkIGFzIEYgdmFsdWVzIHJpZ2h0Pw0KLSBUaGlzIG1lYW5zIHlvdSBjYW4gYWN0dWFsbHkgZ2V0IGFuIEFOT1ZBIGxpa2UgdGFibGUgZm9yIHRoZSBtb2RlbA0KLSBmb3IgZXhhbXBsZTogDQoNCmBgYHtyfQ0KIyMjIyMjIyMjIyMjIyMjTW9kZWwgMSANCkljZS5Nb2RlbDwtbG0oSGFwcGluZXNzfiBJY2VDcmVhbSwgZGF0YSA9IENvcnJEYXRhVCkNCmFub3ZhKEljZS5Nb2RlbCkNCmBgYA0KDQotIFRoZSAkUjIkIHRoaXMgaXMgZXhwbGFpbmVkIHRvIHVuZXhwbGFpbmVkIHZhcmlhbmNlIChsaWtlIGluIG91ciBBTk9WQSkNCi0gJFJeMiA9IFxmcmFje1NTX3tleHBsYWluZWR9fXtTU197ZXhwbGFpbmVkfStTU197cmVzaWR1YWx9fSQNCi0ganVzdCB0byBjaGVjazogYW5vdmEoSWNlLk1vZGVsKSBgciBhbm92YShJY2UuTW9kZWwpJCdTdW0gU3EnWzFdIC8gYW5vdmEoSWNlLk1vZGVsKSQnU3VtIFNxJ1sxXSArIGFub3ZhKEljZS5Nb2RlbCkkJ1N1bSBTcSdbMl1gDQotIHdoaWNoIG1hdGNoZWQgdGhlICRSXjIkIHRoYXQgUiBnaXZlcyB1cyBgciBzdW1tYXJ5KEljZS5Nb2RlbCkkci5zcXVhcmVkYA0KLSBXaGVuIHdlIGNoZWNrIHRvIHNlZSB3aGljaCBtb2RlbCBpcyBiZXN0IHdlIGFjdHVhbGx5IHRlc3QgdGhlIGRpZmZlcmVuY2VzDQoNCiMjIyBMZXRzIGZvcndhcmQtZml0IG91ciBtb2RlbHMNCi0gTW9kZWwgMSAoU21hbGxlciBtb2RlbCkNCg0KYGBge3J9DQpJY2UuTW9kZWw8LWxtKEhhcHBpbmVzc34gSWNlQ3JlYW0sIGRhdGEgPSBDb3JyRGF0YVQpDQpSMi5Nb2RlbC4xPC1zdW1tYXJ5KEljZS5Nb2RlbCkkci5zcXVhcmVkDQpgYGANCg0KLSBNb2RlbCAyIChMYXJnZXIgbW9kZWwpDQoNCmBgYHtyfQ0KIyMjIyMjIyMjIyMjIyMjTW9kZWwgMSANCkljZS5Ccm93bi5Nb2RlbDwtbG0oSGFwcGluZXNzfiBJY2VDcmVhbStCcm93bmllcywgZGF0YSA9IENvcnJEYXRhVCkNClIyLk1vZGVsLjI8LXN1bW1hcnkoSWNlLkJyb3duLk1vZGVsKSRyLnNxdWFyZWQNCmBgYA0KDQoNCmBgYHtyLCByZXN1bHRzPSdhc2lzJ30NCmxpYnJhcnkoc3RhcmdhemVyKQ0Kc3RhcmdhemVyKEljZS5Nb2RlbCxJY2UuQnJvd24uTW9kZWwsdHlwZT0iaHRtbCIsDQogICAgICAgICAgY29sdW1uLmxhYmVscyA9IGMoIk1vZGVsIDEiLCAiTW9kZWwgMiIpLA0KICAgICAgICAgIGludGVyY2VwdC5ib3R0b20gPSBGQUxTRSwNCiAgICAgICAgICBzaW5nbGUucm93PUZBTFNFLCANCiAgICAgICAgICBzdGFyLmN1dG9mZnMgPSBjKDAuMSwgMC4wNSwgMC4wMSwgMC4wMDEpLA0KICAgICAgICAgIHN0YXIuY2hhciA9IGMoIkAiLCAiKiIsICIqKiIsICIqKioiKSwgDQogICAgICAgICAgbm90ZXM9IGMoIkBwIDwgLjEgKnAgPCAuMDUgKipwIDwgLjAxICoqKnAgPCAuMDAxIiksDQogICAgICAgICAgbm90ZXMuYXBwZW5kID0gRkFMU0UsIGhlYWRlcj1GQUxTRSkNCmBgYA0KDQotIExldCdzIHRoZSBkaWZmZXJlbmNlIGluICRSXjIkDQogICAgLSAkUl97Q2hhbmdlfV4yJCA9JFJfe0xhcmdlcn1eMiQgLSAkUl97U21hbGxlcn1eMiQNCi0gSW4gUiwgd2UgY2FsbCBmb3IgZnVuY3Rpb24gYGFub3ZhYCBhbmQgdXNlIGFuICRGJCB3aGVyZSB0aGUgZGVncmVlcyBvZiBmcmVlZG9tIGlzIHRoZSBudW1iZXIgb2YgcGFyYW1ldGVyIGRpZmZlcmVuY2VzIGJldHdlZW4gTGFyZ2VyIGFuZCBTbWFsbGVyIG1vZGVsDQoNCmBgYHtyLCBlY2hvPVRSVUUsIHdhcm5pbmc9RkFMU0V9DQpSMi5DaGFuZ2U8LVIyLk1vZGVsLjItUjIuTW9kZWwuMQ0KYW5vdmEoSWNlLk1vZGVsLEljZS5Ccm93bi5Nb2RlbCkNCmBgYA0KDQotIFRoZSAkUl97Q2hhbmdlfV4yJCA9IGByIFIyLkNoYW5nZWAgaXMgc2lnbmlmaWNhbnQgIA0KLSBTbywgaW4gb3RoZXIgd29yZHMsIHdlIHNlZSBtb2RlbCAyICpmaXQqIHRoZSBkYXRhIGJldHRlciB0aGFuIG1vZGVsIDEuIA0KDQoNCiMjIyBCYWNrd2FyZC1maXR0aW5nIG9mIG5lc3RlZCBtb2RlbHMNCi0gWW91IGFzIGRvZXMgdGFraW5nIGF3YXkgdmFyaWFibGVzIHJlZHVjZSBteSAkUl4yJCBzaWduaWZpY2FudGx5IA0KLSBTb21ldGltZXMgdXNlZCB0byB2YWxpZGF0ZSB5b3UgaGF2ZSBhIHBhcnNpbW9uaW91cyBtb2RlbA0KLSBZb3UgbWlnaHQgZm9yd2FyZC1maXQgYSAqc2V0KiBvZiB2YXJpYWJsZXMgYW5kIGJhY2t3YXJkIGZpdCBjcml0aWNhbCBvbmVzIHRvIHRlc3QgYSBzcGVjaWZpYyBoeXBvdGhlc2lzDQotIFVzaW5nIHRoZSBzYW1lIGRhdGEgYXMgYWJvdmUsIHdlIHdpbGwgZ2V0IHRoZSBzYW1lIHZhbHVlcyAoanVzdCBuZWdhdGl2ZSkNCiAgICAtICRSX3tDaGFuZ2V9XjIkID0kUl97c21hbGxlcn1eMiQgLSAkUl97TGFyZ2VyfV4yJA0KDQpgYGB7cn0NCiMjIyMjIyMjIyMjIyMjI01vZGVsIDEuQiANCkljZS5Ccm93bi5Nb2RlbDwtbG0oSGFwcGluZXNzfiBJY2VDcmVhbStCcm93bmllcywgZGF0YSA9IENvcnJEYXRhVCkNClIyLk1vZGVsLjEuQjwtc3VtbWFyeShJY2UuQnJvd24uTW9kZWwpJHIuc3F1YXJlZA0KIyMjIyMjIyMjIyMjIyMjTW9kZWwgMi5CDQpJY2UuTW9kZWw8LWxtKEhhcHBpbmVzc34gSWNlQ3JlYW0sIGRhdGEgPSBDb3JyRGF0YVQpDQpSMi5Nb2RlbC4yLkI8LXN1bW1hcnkoSWNlLk1vZGVsKSRyLnNxdWFyZWQNClIyLkNoYW5nZS5CPC1SMi5Nb2RlbC4yLkItUjIuTW9kZWwuMS5CDQphbm92YShJY2UuQnJvd24uTW9kZWwsSWNlLk1vZGVsKQ0KYGBgDQoNCi0gVGhlICRSX3tDaGFuZ2V9XjIkID0gYHIgUjIuQ2hhbmdlLkJgIGlzIHNpZ25pZmljYW50ICANCi0gU28sIGluIG90aGVyIHdvcmRzLCB3ZSBzZWUgbW9kZWwgMSBpcyBhIHdvcnNlIGZpdCBvZiB0aGUgZGF0YSB0aGFuIG1vZGVsIDIgDQoNCg0KIyMgU3RlcHdpc2UgbW9kZWxpbmcgYnkgQ29tcHV0ZXINCi0gU3RlcHdpc2Ugd2l0aCBtYW55IHByZWRpY3RzIGlzIG9mdGVuIGRvbmUgYnkgY29tcHV0ZXIgYW5kIGl0IGRvZXMgbm90IGFsd2F5cyBhc3N1bWUgbmVzdGVkIG1vZGVscyAoeW91IGNhbiBhZGQgYW5kIHJlbW92ZSBhdCB0aGUgc2FtZSkNCi0gRXhwbG9yYXRvcnk6IHlvdSBoYXZlIHRvbyBtYW55IHByZWRpY3RvcnMgYW5kIGhhdmUgbm8gaWRlYSB3aGVyZSB0byBzdGFydA0KLSBZb3UgZ2l2ZSB0aGUgY29tcHV0ZXIgYSBsYXJnZXIgbnVtYmVyIG9mIHByZWRpY3RvcnMsIGFuZCB0aGUgY29tcHV0ZXIgZGVjaWRlcyB0aGUgYmVzdCBmaXQgbW9kZWwNCi0gU291bmRzIGdvb2QsIHJpZ2h0PyBObywgYXMgdGhlIHJlc3VsdHMgY2FuIGJlIHVuc3RhYmxlDQogICAgLSBDaGFuZ2Ugb25lIHZhcmlhYmxlIGluIHRoZSBzZXQgYW5kIHRoZSBmaW5hbCBtb2RlbCBjYW4gY2hhbmdlDQogICAgLSBIaWdoIGNoYW5jZSBvZiB0eXBlIEkgYW5kIHR5cGUgSUkgZXJyb3INCiAgICAtIFRoZSBjb21wdXRlciBtYWtlcyBkZWNpc2lvbnMgYmFzZWQgb24gQWthaWtlIGluZm9ybWF0aW9uIGNyaXRlcmlvbiAoQUlDKSBub3Qgc2VsZWN0ZWQgYmFzZWQgb24gYSBjaGFuZ2UgaW4gJFJeMiQsIGJlY2F1c2UgbW9kZWxzIGFyZSBub3QgbmVzdGVkDQogICAgLSBhbHNvIGNvbXB1dGVyIG1ha2VzIGRlY2lzaW9ucyBwdXJlbHkgb24gZml0IHZhbHVlcyBhbmQgaGFzIG5vdGhpbmcgZG8gd2l0aCBhIHRoZW9yeQ0KICAgIC0gU29sdXRpb25zIGFyZSBvZnRlbiB1bmlxdWUgdG8gdGhhdCBwYXJ0aWN1bGFyIGRhdGFzZXQNCiAgICAtIFRoZSBiZXN0IG1vZGVsIGlzIG9mdGVuIHRoZSBvbmUgdGhhdCBwYXJzZXMgYSB0aGVvcnkgYW5kIG9ubHkgYSBodW1hbiBjYW4gZG8gdGhhdCBhdCBwcmVzZW50DQotIE5vdCByZWFsbHkgcHVibGlzaGFibGUgYmVjYXVzZSBvZiB0aGVzZSBwcm9ibGVtcw0KDQojIFBhcnNpbmcgaW5mbHVlbmNlDQotIEFzIG1vZGVscyBnZXQgYmlnZ2VyIGFuZCBiaWdnZXIgaXRzIGJlY29tZXMgYSBjaGFsbGVuZ2UgdG8gZmlndXJlIG91dCB0aGUgdW5pcXVlIGNvbnRyaWJ1dGlvbiB0byAkUl4yJCBvZiBlYWNoIHZhcmlhYmxlDQotIFRoZXJlIGFyZSBtYW55IGNvbXB1dGF0aW9uIHNvbHV0aW9ucyB0aGF0IHlvdSBjYW4gc2VsZWN0IGZyb20sIGJ1dCB3ZSB3aWxsIHVzZSBvbmUgY2FsbGVkICoqbG1nKioNCi0geW91IGNhbiByZWFkIGFib3V0IGFsbCB0aGUgZGlmZmVyZW50IG9uZXMgaGVyZTogPGh0dHBzOi8vY29yZS5hYy51ay9kb3dubG9hZC9wZGYvNjMwNTAwNi5wZGY+DQotIHRoZXNlIG1ldGhvZHMgYXJlIG5vdCB3ZWxsIGtub3duIGluIHBzeWNob2xvZ3ksIGJ1dCBjYW4gYmUgdmVyeSB1c2VmdWwgd2hlbiBwZW9wbGUgYXNrIHlvdSB3aGF0IHRoZSByZWxhdGl2ZSBpbXBvcnRhbmNlIG9mIGVhY2ggdmFyaWFibGUgaXMNCi0gdHdvIGFwcHJvYWNoZXM6IHNob3cgYWJzb2x1dGUgJFJeMiQgZm9yIGVhY2ggdGVybSBvciB0aGUgcmVsYXRpdmUgJSBvZiAkUl4yJCBmb3IgZWFjaCB0ZXJtDQoNCmBgYHtyLCBlY2hvPVRSVUUsIHdhcm5pbmc9RkFMU0UsIG1lc3NhZ2U9RkFMU0V9DQpsaWJyYXJ5KHJlbGFpbXBvKQ0KIyBJbiB0ZXJtcyBvZiBSMg0KY2FsYy5yZWxpbXAoSWNlLkJyb3duLk1vZGVsKSANCiMgYXMgJSBvZiBSMg0KY2FsYy5yZWxpbXAoSWNlLkJyb3duLk1vZGVsLHJlbGEgPSBUUlVFKSANCmBgYA0KDQoNCiMgRmluYWwgbm90ZXM6IA0KLSBJZiB5b3UgcGxheSB3aXRoIGxvdHMgb2YgcHJlZGljdG9ycyBhbmQgZG8gbG90cyBvZiBtb2RlbHMsIHNvbWV0aGluZyB3aWxsIGJlIHNpZ25pZmljYW50DQotIFR5cGUgSSBlcnJvciBpcyBhIGJpZyBwcm9ibGVtIGJlY2F1c2Ugb2YgdGhlICdyZXNlYXJjaGVyIGRlZ3JlZSBvZiBmcmVlZG9tIHByb2JsZW0nDQotIFR5cGUgSUkgaW5jcmVhc2VzIGFzIGEgZnVuY3Rpb24gb2YgdGhlIG51bWJlciBvZiBwcmVkaWN0b3JzLiBhKSB5b3Ugc2xpY2UgdG9vIG11Y2ggcGllLCBiKSBlYWNoIHZhcmlhYmxlIG1pZ2h0IHRyeSB0byBlYWNoIGVhdCBzb21lb25lIGVsc2UncyBzbGljZQ0KLSBMZXNzIGlzIG1vcmU6IGFzayB0YXJnZXRlZCBxdWVzdGlvbnMgd2l0aCBhcyBvcnRob2dvbmFsIGEgc2V0IG9mIHZhcmlhYmxlcyBhcyB5b3UgY2FuIA0KPHNjcmlwdD4NCiAgKGZ1bmN0aW9uKGkscyxvLGcscixhLG0pe2lbJ0dvb2dsZUFuYWx5dGljc09iamVjdCddPXI7aVtyXT1pW3JdfHxmdW5jdGlvbigpew0KICAoaVtyXS5xPWlbcl0ucXx8W10pLnB1c2goYXJndW1lbnRzKX0saVtyXS5sPTEqbmV3IERhdGUoKTthPXMuY3JlYXRlRWxlbWVudChvKSwNCiAgbT1zLmdldEVsZW1lbnRzQnlUYWdOYW1lKG8pWzBdO2EuYXN5bmM9MTthLnNyYz1nO20ucGFyZW50Tm9kZS5pbnNlcnRCZWZvcmUoYSxtKQ0KICB9KSh3aW5kb3csZG9jdW1lbnQsJ3NjcmlwdCcsJ2h0dHBzOi8vd3d3Lmdvb2dsZS1hbmFseXRpY3MuY29tL2FuYWx5dGljcy5qcycsJ2dhJyk7DQoNCiAgZ2EoJ2NyZWF0ZScsICdVQS05MDQxNTE2MC0xJywgJ2F1dG8nKTsNCiAgZ2EoJ3NlbmQnLCAncGFnZXZpZXcnKTsNCg0KPC9zY3JpcHQ+DQo=